Data Wrangling
Grayson White
Math 241
Week 3 | Spring 2026
Announcements
- Office Hours Schedule
- Problem Set 1 due Thursday 9am.
Week 3 Goals
Mon Lecture
Think about reproducibility and learn how to ask coding questions well.
Motivate and work on data wrangling.
Wed Lecture
- Plot animation and interactivity.
- GitHub workflow review.
Reproducible Workflow
One where if you shared your data and work with someone else, they could reproduce your results.
Not the same as replication: Where someone collects new data following your same design to see if they get the same results.
Quartodocuments allow us to include ourRcode, output, and narrative in the same place.- Load the raw data.
- Be transparent about all the analysis steps.
- Even if you don’t showcase the
Rcode in the output file, it is contained in theqmdfile.
Creating reproducible examples with reprex

Why do I need to learn to create reproducible technical examples?
- So that you can ask and answer questions in our class Slack Workspace or Stack Overflow or other
Rhelp sites!
First, let’s take a look at some bad examples!
What is wrong with this coding question?
I am trying to create a plot and I can’t get the bars to do what I want them to. Help?!
What is wrong with this coding question?
I want to do the following but it isn’t working:
thing <- read.csv(“long/file/path/thing.csv”)
ggplot(thing, aes(x = factor(that))) + geom_bar()
Help?!
What is wrong with this coding question?
I want to reorder the bars of my plot but can’t get it working. Help!
library(tidyverse)
library(palmerpenguins)
penguins <- penguins %>%
group_by(species) %>%
mutate(mean_flipper = mean(flipper_length_mm)) %>%
ungroup() %>%
mutate(long = case_when(flipper_length_mm < mean(flipper_length_mm) ~ "no",
flipper_length_mm >= mean(flipper_length_mm) ~ "yes"))
penguins %>%
ggplot(mapping = aes(x = factor(species))) +
geom_bar()What is wrong with this coding question?
I want to reorder the bars of my plot but can’t get it working. Help!
What makes a good coding question?
It uses a minimal dataset to reproduce the issue.
It includes the shortest amount of runnable code necessary to reproduce the issue.
It doesn’t wreak havoc on other people’s computers.
It includes code and output so that others don’t have to run it!
- It includes any necessary information on the used packages, R version, system, etc.
- Can use
packageVersion("tidyverse")orsessionInfo()to find this information.
- Can use
Minimal Dataset: two good options
Create a toy data frame.
Use a built-in dataset or a dataset from a particular package.
# A tibble: 344 × 10
species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
<fct> <fct> <dbl> <dbl> <int> <int>
1 Adelie Torgersen 39.1 18.7 181 3750
2 Adelie Torgersen 39.5 17.4 186 3800
3 Adelie Torgersen 40.3 18 195 3250
4 Adelie Torgersen NA NA NA NA
5 Adelie Torgersen 36.7 19.3 193 3450
6 Adelie Torgersen 39.3 20.6 190 3650
7 Adelie Torgersen 38.9 17.8 181 3625
8 Adelie Torgersen 39.2 19.6 195 4675
9 Adelie Torgersen 34.1 18.1 193 3475
10 Adelie Torgersen 42 20.2 190 4250
# ℹ 334 more rows
# ℹ 4 more variables: sex <fct>, year <int>, mean_flipper <dbl>, long <chr>Minimal Code
Include the necessary libraries.
Test run the code in a restarted R session to make sure it is runnable!
Make sure your code is copy-and-paste-able!
Don’t copy from the console.
Make sure your code is copy-and-paste-able!

Now we have our reproducible example:
How can I reorder the bars in the ggplot to go from the most frequent to the least frequent category?
How can we easily share it?
- Using the
reprex()function in thereprexpackage.
reprex Practice Time!
But first: Q: What is an R script file?
- A text file for entering R commands.
Q: How is an R script file different from a Quarto or RMarkdown document?
- You only put code in an
Rscript.
- If you add any text you must comment it out with
#. - Think of it as a single
Rchunk that you won’t knit into an output document. - Useful when writing a lot of code and want to compartmentalize.
reprex Practice Time!
In Session, select “Clear Workspace” and then “Restart R”.
Open a script file and include in the top line:
- Put the code you want to use in the script file and make sure it runs.
Surround the code with
reprex({ ... }, venue = "slack")and run it.An md file will pop up. Copy all the contents of that file.
Head over to the
#coding-qachannel and paste in the contents as a reply to thereprexpractice message. A text box will pop-up and select “Apply”.Above your code, type your question. Then hit “Send”.
Data Wrangling
Data Wrangling
- So far, we’ve focused primarily on data visualization!
- However, in order to make beautiful graphics, sometimes we have to go from
messy data ➡️ beautiful, wrangled, and cleaned data
Example: Data In the Wild
Many datasets on the internet are often in Display Format, not Analysis Format.
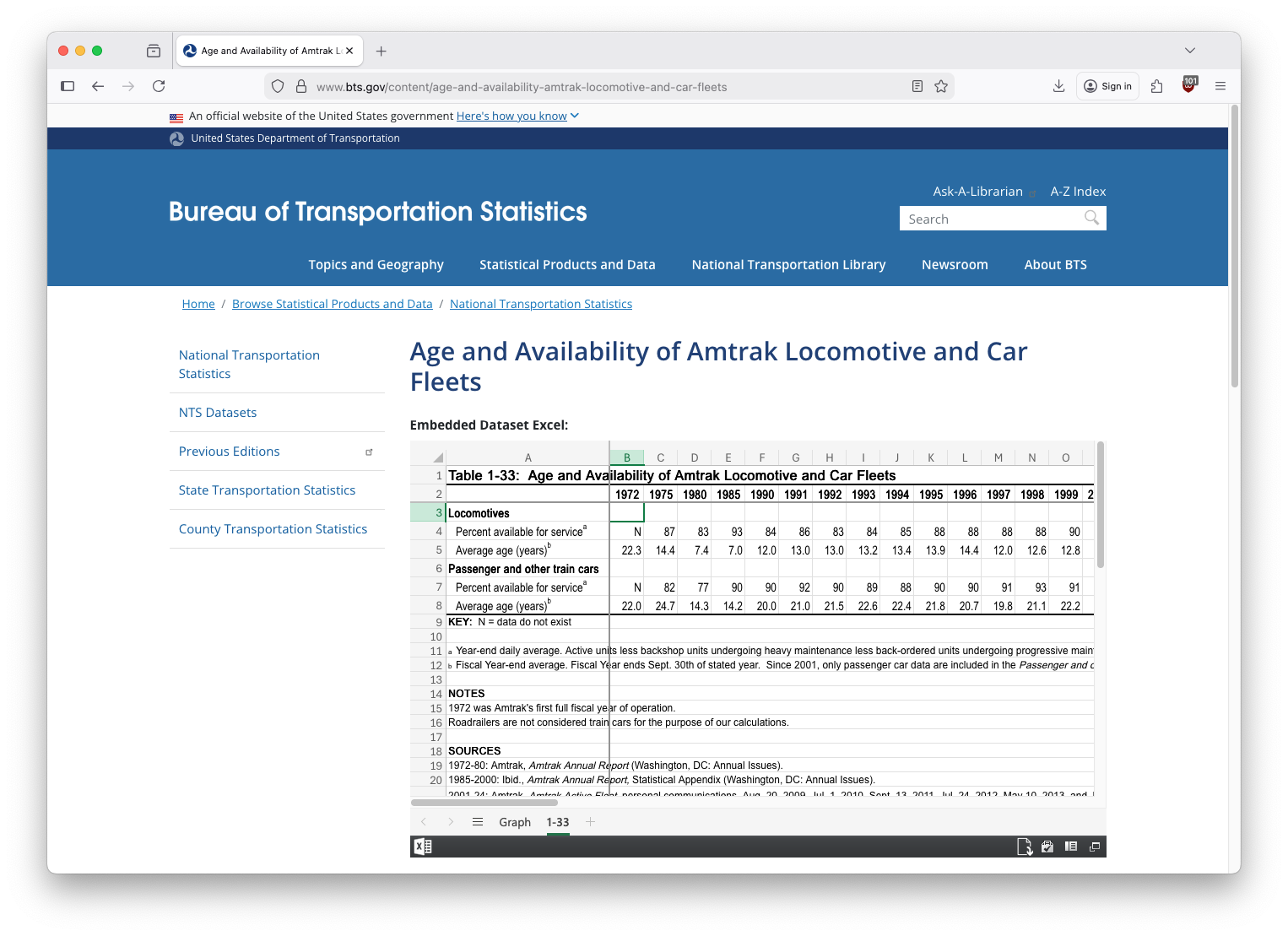
And, sometimes we’d like to modify a dataset to help us answer a specific question, or display the data in a particular way.
Many of you have already needed to do some data wrangling in Problem Set 1.
Recall this visualization from Week 1, Day 1
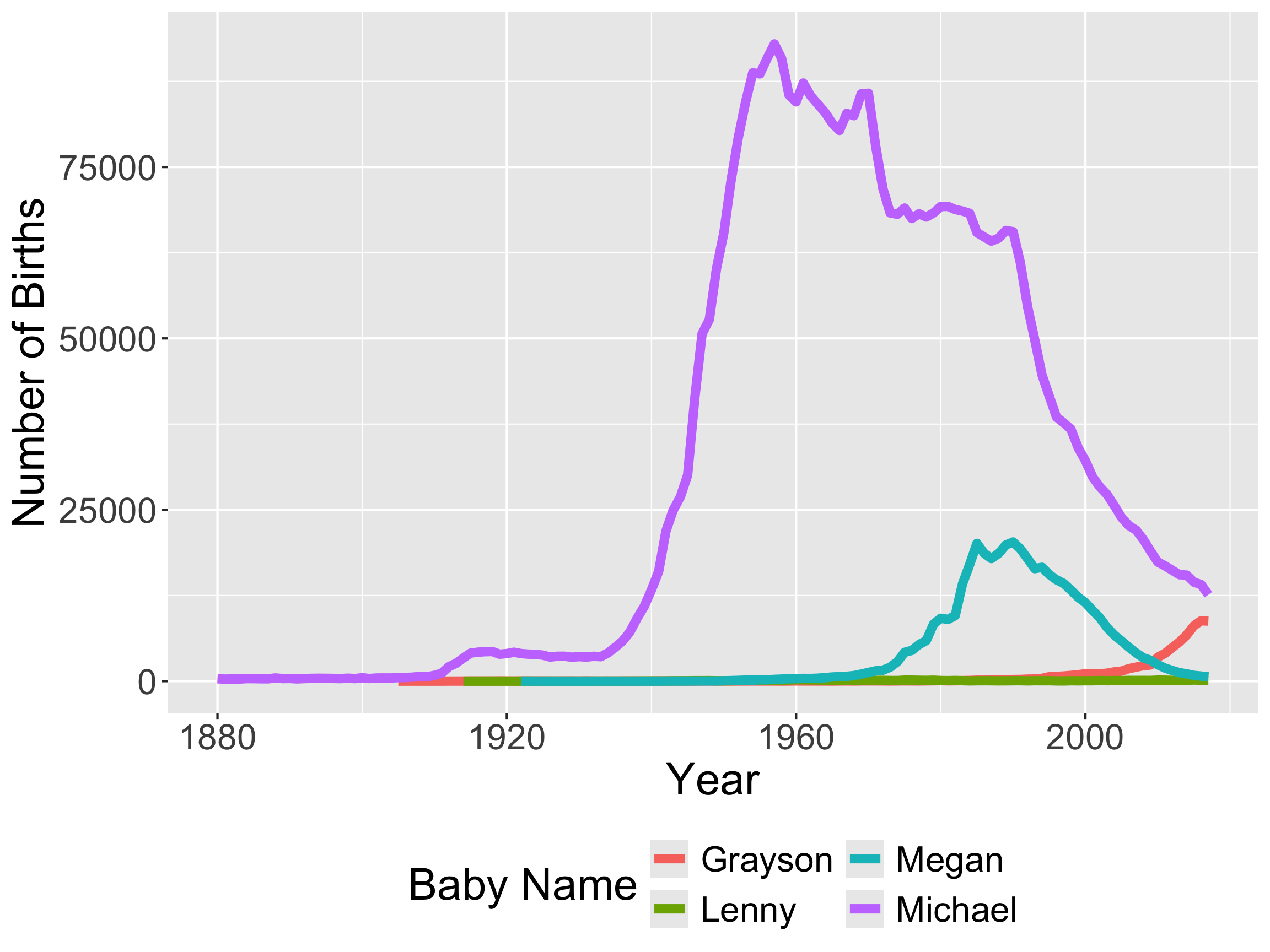
- We need to be able to wrangle the data to make this visualization!
Recall this visualization from Week 1, Day 1
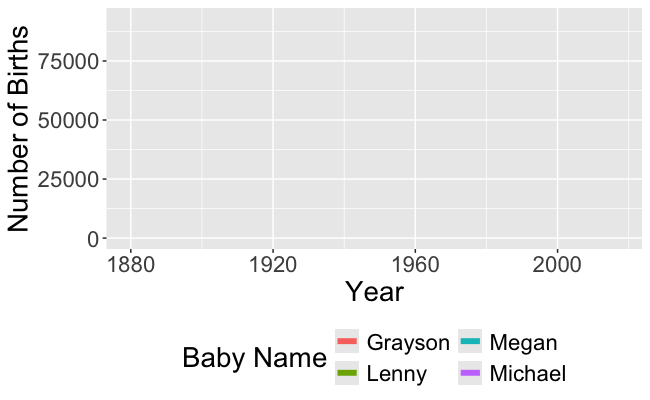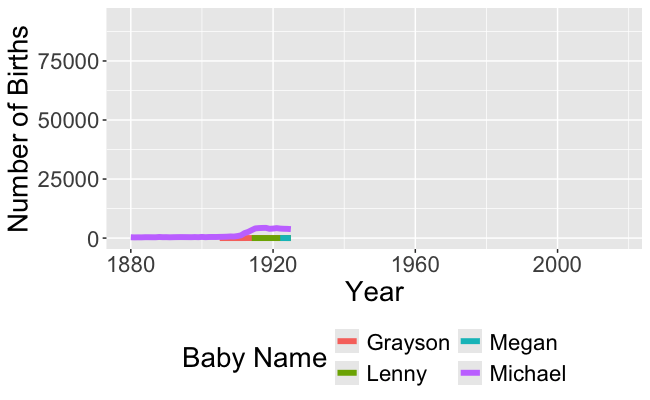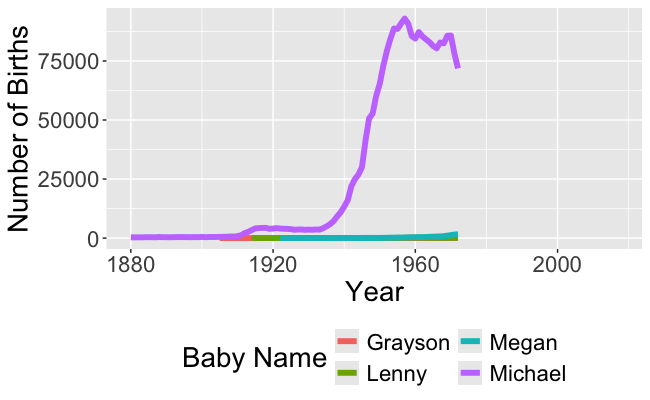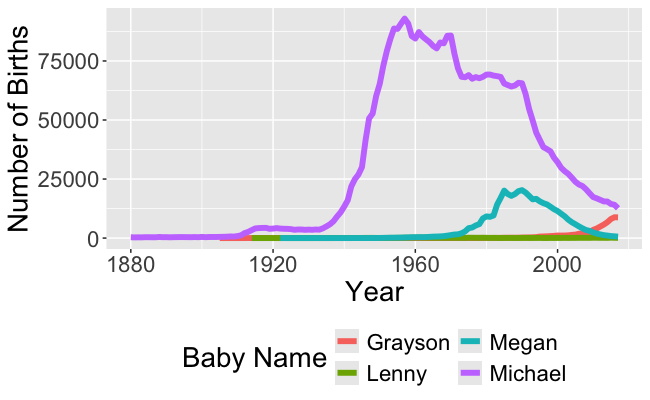
- And to make the animated version!
- In order to make these visualizations, we have to learn how to wrangle data!
Data Wrangling
- What is it?
- Any processing you have to do to the data to summarize, visualize, model it.
- Examples:
- Recoding 999 as “NA”
- Removing rows
- Creating new variables
- Recoding category variables
- Fixing variable types
- Reshaping data into a format that satisfies the tidy data principles
Data Wrangling
We are going to learn several packages to help us wrangle data.
We’ll learn the core of
dplyrright now!

Math 241 dplyr Experience
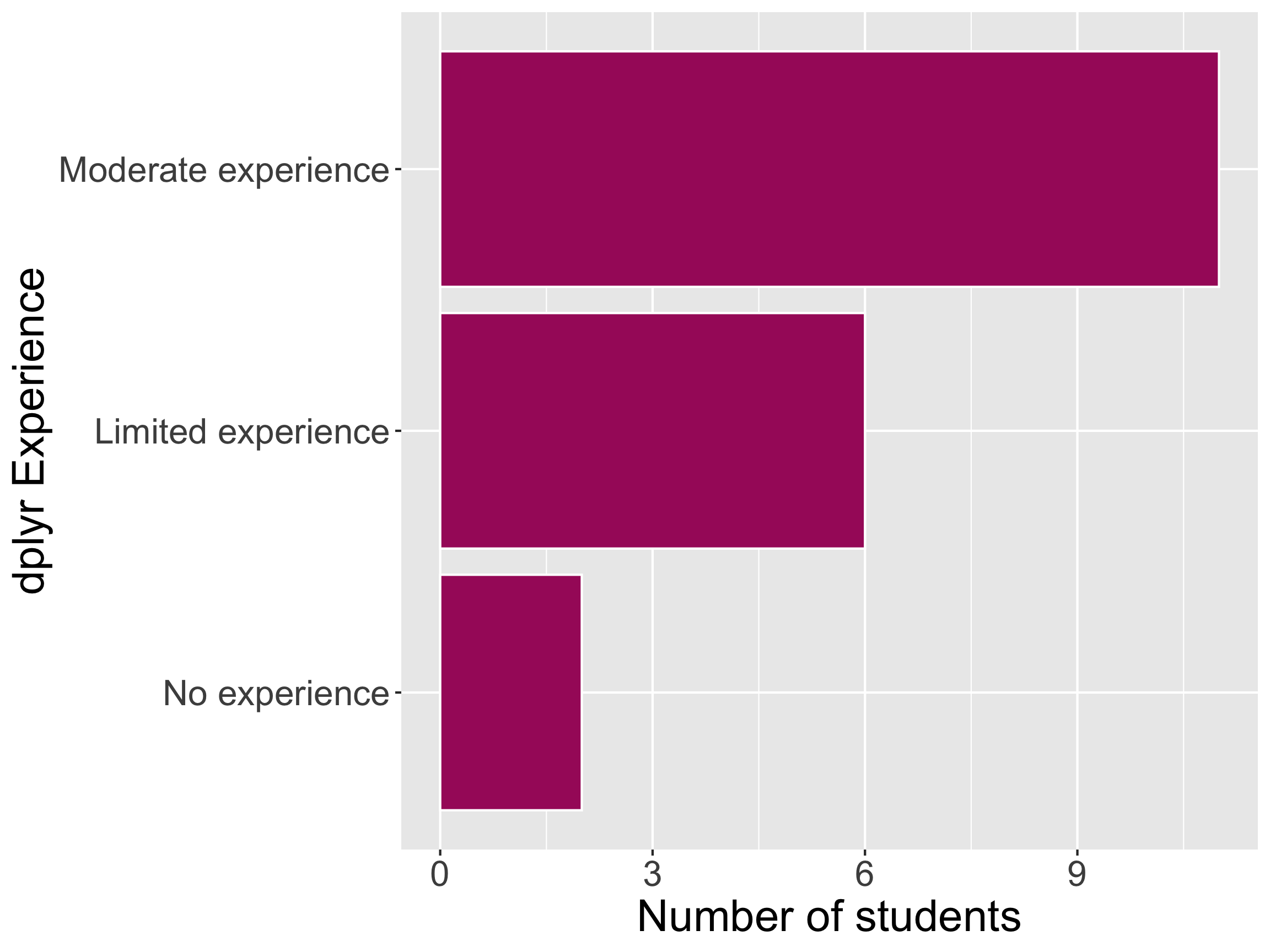
dplyr core functions
dplyr for Data Wrangling
- Common wrangling verbs:
select()mutate()filter()arrange()summarize()drop_na()count()
- One action:
group_by()
Pipe for chaining together commands
magrittrpipe:%>%- New base
R(native) pipe:|>- Less flexible in some scenarios, but…
- Don’t need to load a package to use it
New data: babynames
# A tibble: 6 × 5
year sex name n prop
<dbl> <chr> <chr> <int> <dbl>
1 1880 F Mary 7065 0.0724
2 1880 F Anna 2604 0.0267
3 1880 F Emma 2003 0.0205
4 1880 F Elizabeth 1939 0.0199
5 1880 F Minnie 1746 0.0179
6 1880 F Margaret 1578 0.0162- What does each row represent?
Wrangling babynames
What wrangling do we need to do to get from raw data to wrangled data?
Wrangling babynames
filter(): Subset rows based on logic statements.
# A tibble: 675 × 5
year sex name n prop
<dbl> <chr> <chr> <int> <dbl>
1 1880 M Michael 354 0.00299
2 1881 M Michael 298 0.00275
3 1882 M Michael 321 0.00263
4 1883 M Michael 307 0.00273
5 1884 M Michael 373 0.00304
6 1885 M Michael 370 0.00319
7 1886 M Michael 348 0.00292
8 1887 M Michael 345 0.00316
9 1888 M Michael 466 0.00359
10 1889 M Michael 377 0.00317
# ℹ 665 more rowsWrangling babynames
%>% (the pipe): Inserts current line as first argument in the next line.
# A tibble: 675 × 5
year sex name n prop
<dbl> <chr> <chr> <int> <dbl>
1 1880 M Michael 354 0.00299
2 1881 M Michael 298 0.00275
3 1882 M Michael 321 0.00263
4 1883 M Michael 307 0.00273
5 1884 M Michael 373 0.00304
6 1885 M Michael 370 0.00319
7 1886 M Michael 348 0.00292
8 1887 M Michael 345 0.00316
9 1888 M Michael 466 0.00359
10 1889 M Michael 377 0.00317
# ℹ 665 more rowsWrangling babynames
%>% (the pipe): Inserts current line as first argument in the next line.
# A tibble: 675 × 5
year sex name n prop
<dbl> <chr> <chr> <int> <dbl>
1 1880 M Michael 354 0.00299
2 1881 M Michael 298 0.00275
3 1882 M Michael 321 0.00263
4 1883 M Michael 307 0.00273
5 1884 M Michael 373 0.00304
6 1885 M Michael 370 0.00319
7 1886 M Michael 348 0.00292
8 1887 M Michael 345 0.00316
9 1888 M Michael 466 0.00359
10 1889 M Michael 377 0.00317
# ℹ 665 more rowsWrangling babynames
summarize(): Compute an aggregation measure (e.g., mean, standard deviation).
Wrangling babynames
group_by(): Create groups for how future operations should be computed.
# A tibble: 4 × 2
name n
<chr> <int>
1 Grayson 74707
2 Lenny 7768
3 Megan 437623
4 Michael 4372536- What are we missing?
Wrangling babynames
group_by(): Create groups for how future operations should be computed.
# A tibble: 431 × 3
# Groups: year [138]
year name n
<dbl> <chr> <int>
1 1880 Michael 354
2 1881 Michael 298
3 1882 Michael 321
4 1883 Michael 307
5 1884 Michael 373
6 1885 Michael 370
7 1886 Michael 348
8 1887 Michael 345
9 1888 Michael 466
10 1889 Michael 377
# ℹ 421 more rowsWrangling babynames
group_by(): Create groups for how future operations should be computed.
# A tibble: 431 × 3
year name n
<dbl> <chr> <int>
1 1880 Michael 354
2 1881 Michael 298
3 1882 Michael 321
4 1883 Michael 307
5 1884 Michael 373
6 1885 Michael 370
7 1886 Michael 348
8 1887 Michael 345
9 1888 Michael 466
10 1889 Michael 377
# ℹ 421 more rows- What did
ungroup()do?
Wrangling babynames
arrange(): Order the rows by variable(s).
# A tibble: 431 × 3
year name n
<dbl> <chr> <int>
1 2017 Grayson 8767
2 2017 Lenny 118
3 2017 Megan 624
4 2017 Michael 12612
5 2016 Grayson 8817
6 2016 Lenny 134
7 2016 Megan 744
8 2016 Michael 14091
9 2015 Grayson 8071
10 2015 Lenny 177
# ℹ 421 more rowsNew data: pets I’ve had
Wrangling my_pets
select(): retain particular columns.
Wrangling my_pets
select(): retain particular columns.
Wrangling my_pets
drop_na(): Remove any row that has a missing value for particular variables.
Wrangling my_pets
mutate(): Create new variables or modify existing variables
- Use case: pet years!
- 8 for large dog, 7 for cats
- (I’m not an animal scientist)
Wrangling my_pets
mutate(): Create new variables or modify existing variables
- Use case: pet years!
- 8 for large dog, 7 for cats
- (I’m not an animal scientist)
Better solution: use case_when()
Wrangling my_pets
More filter()ing
Wrangling my_pets
More filter()ing: Nubs seemed like he was pretty old
[1] name type age birthplace age_pet_years
<0 rows> (or 0-length row.names)What went wrong here???
Logical operators:
&means ‘and’
Wrangling my_pets
More filter()ing: Nubs seemed like he was pretty old
name type age birthplace age_pet_years
1 Dude dog 8 Washington 64
2 Pickle cat 6 <NA> 42
3 Kyle cat 4 Utah 28
4 Nubs cat NA <NA> NA- Logical operators:
|means ‘or’
Wrangling my_pets
More filter()ing
name type age birthplace age_pet_years
1 Dude dog 8 Washington 64- In a
filter(), the comma implicitly means&
Wrangling my_pets
%in% vs. ==
name type age birthplace age_pet_years
1 Dude dog 8 Washington 64- What went wrong here?
Wrangling my_pets
%in% vs. ==
Activity
15:00
Next time
- Use some of our data wrangling skills to make animated and interactive graphics!